-Search query
-Search result
Showing all 29 items for (author: olson & wc)

EMDB-41888: 
Structure of Apo CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41889: 
Structure of CXCL12-bound CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41890: 
Structure of AMD3100-bound CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41891: 
Structure of REGN7663 Fab-bound CXCR4/Gi complex
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41892: 
Structure of REGN7663-Fab bound CXCR4
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41893: 
Structure of trimeric CXCR4 in complex with REGN7663 Fab
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-41894: 
Structure of tetrameric CXCR4 in complex with REGN7663 Fab
Method: single particle / : Saotome K, McGoldrick LL, Franklin MC

EMDB-28570: 
CryoEM structure of PN45545 TCR-CD3 complex
Method: single particle / : Saotome K, Franklin MC

EMDB-28571: 
CryoEM structure of PN45545 TCR-CD3 in complex with HLA-A2 MAGEA4 (230-239)
Method: single particle / : Saotome K, Franklin MC

EMDB-28572: 
CryoEM structure of PN45428 TCR-CD3 in complex with HLA-A2 MAGEA4
Method: single particle / : Saotome K, Franklin MC

EMDB-28573: 
CryoEM structure of HLA-A2 bound to MAGEA4 (230-239) peptide
Method: single particle / : Saotome K, Franklin MC

EMDB-28574: 
CryoEM structure of HLA-A2 bound to MAGEA8 (232-241) peptide
Method: single particle / : Saotome K, Franklin MC

EMDB-27221: 
Cryo-EM map of human LIF signaling complex with full extracellular domains
Method: single particle / : Zhou Y, Franklin MC

EMDB-27227: 
Cryo-EM map of human CNTF signaling complex with full extracellular domains: refined from selected full particle dataset, with better resolution in the interaction core region
Method: single particle / : Zhou Y, Franklin MC

EMDB-27228: 
Cryo-EM structure of human CNTFR alpha in complex with the Fab fragments of two antibodies
Method: single particle / : Zhou Y, Franklin MC

EMDB-27229: 
Cryo-EM map of human CNTF signaling complex with full extracellular domains: refined from a particle subset, with lower global resolution but improved LIFR D6-D8 density
Method: single particle / : Zhou Y, Franklin MC

EMDB-27230: 
Cryo-EM structure of human CLCF1 in complex with CRLF1 and CNTFR alpha
Method: single particle / : Zhou Y, Franklin MC

EMDB-27231: 
Cryo-EM map of human CLCF1 signaling complex with full extracellular domains
Method: single particle / : Zhou Y, Franklin MC

EMDB-27244: 
Cryo-EM map of detergent-solubilized human IL-6 signaling complex with full extracellular domains
Method: single particle / : Zhou Y, Franklin MC

EMDB-27246: 
Cryo-EM map of human IL-27 signaling complex with full extracellular domains
Method: single particle / : Zhou Y, Franklin MC

EMDB-27247: 
Cryo-EM map of human IL-27 signaling complex: focused refinement on the interaction core region
Method: single particle / : Zhou Y, Franklin MC

EMDB-1930: 
Molecular structure of soluble trimeric HIV-1 glycoprotein gp140 JR-FL
Method: single particle / : Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S

EMDB-5320: 
Molecular structure of soluble trimeric HIV-1 glycoprotein gp140 KNH1144
Method: subtomogram averaging / : Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S
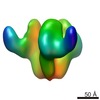
EMDB-5321: 
Molecular structure of soluble trimeric HIV-1 glycoprotein gp140 KNH1144 with sCD4
Method: subtomogram averaging / : Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S

EMDB-5322: 
Molecular structure of soluble trimeric HIV-1 glycoprotein gp140 JR-FL with sCD4
Method: subtomogram averaging / : Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S

EMDB-5323: 
Molecular structure of soluble trimeric HIV-1 glycoprotein gp140 KNH1144 with sCD4 and 17b
Method: subtomogram averaging / : Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S

EMDB-5324: 
Molecular structure of soluble trimeric HIV-1 glycoprotein gp140 JR-FL with sCD4 and 17b
Method: subtomogram averaging / : Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S

EMDB-5325: 
Molecular structure of soluble trimeric HIV-1 glycoprotein gp140 KNH1144 with 17b
Method: subtomogram averaging / : Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
